YGR273C Yeast ORFan Analysis
About Yeast ORFan Gene Project
The Yeast ORFan Gene Project is a consortium of undergraduate researchers and faculty at primarily undergraduate institutions (PUIs) to coordinate resources and design strategies to assign molecular functions to genes of unknown function in the model organism S. cerevisiae (Baker’s yeast). For more information, please visit here.
About YGR273C
YGR273C is a putative protein of unknown function. More information about YGR273C can be found in the Saccharomyces Genome Database.
Summary of Findings for YGR273C
This project aims to uncover clues of uncharacterized Saccharomyces Cerevisiae gene YGR273C using existing data without performing experiments. Current data suggests that YGR273C is a non-essential gene involved in the later stages of sporulation. It localizes to the cytosol and possibly promotes filament growth during spore wall formation.
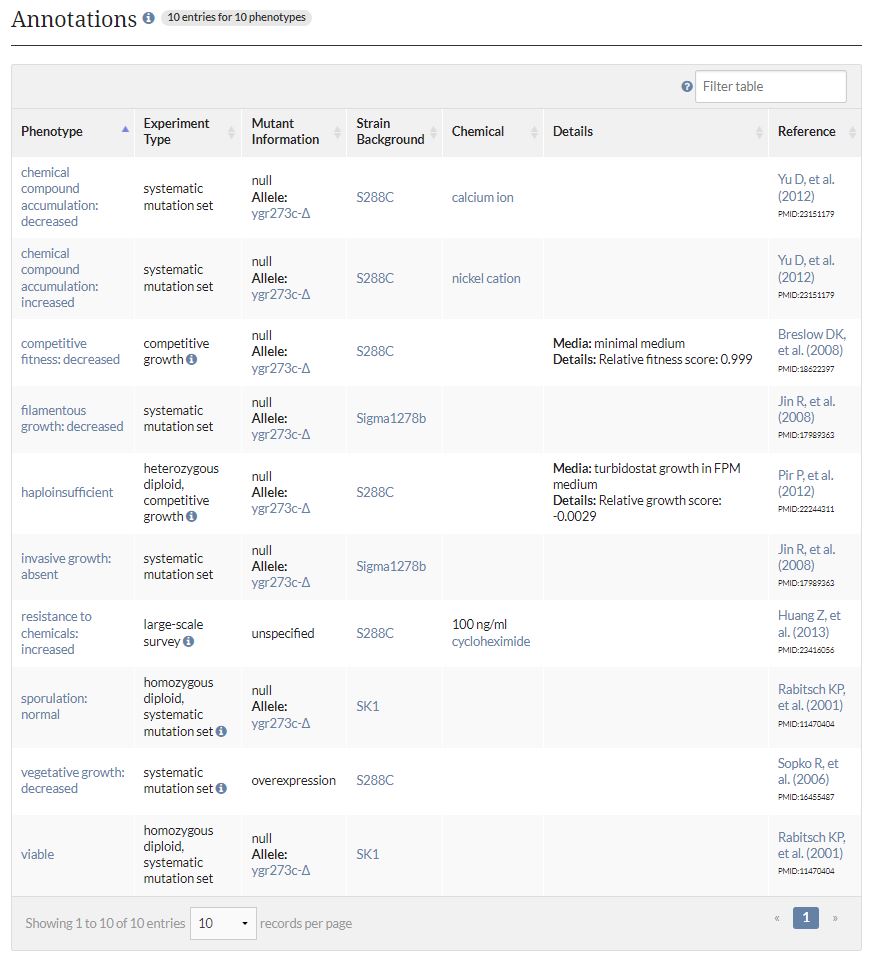
Diploid Saccharomyces Cerevisiae cells undergo sporulation in response to nutritional stress, forming thick spore walls to withstand environmental stress. The deletion of YGR273C does not affect normal sporulation but results in haploid insufficient expression, a decrease in filamentous growth, and reduced cell competitiveness. The YGR273C null mutation test results suggest that YGR273C possibly promotes filamentous growth, contributing to the formation of rigid spore walls. Click on the image below to see null mutation test results.
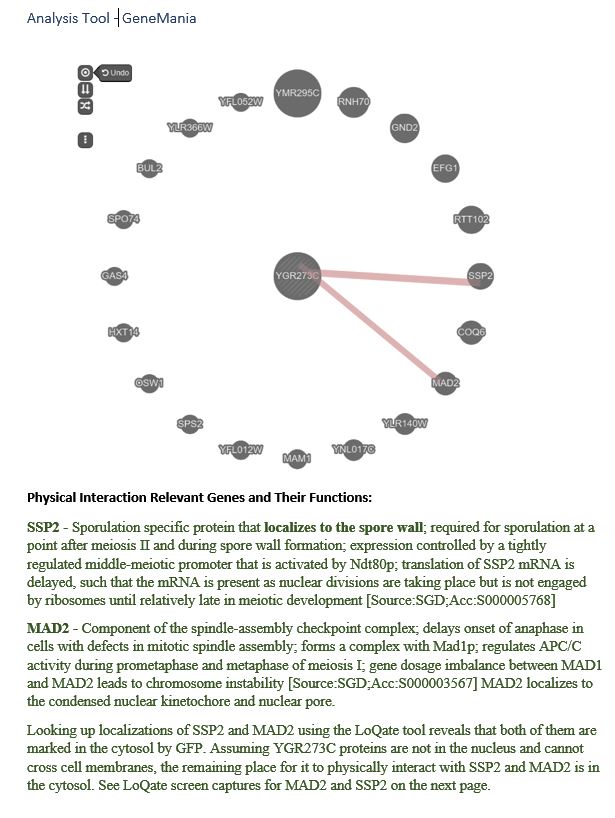
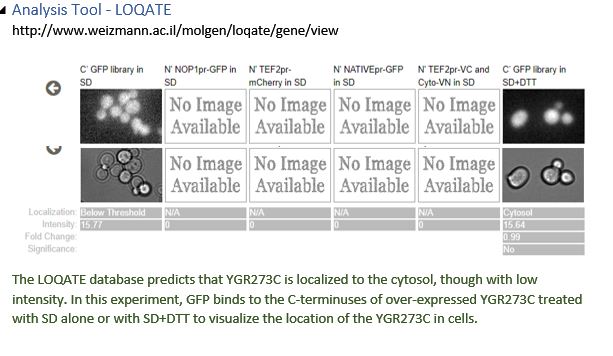
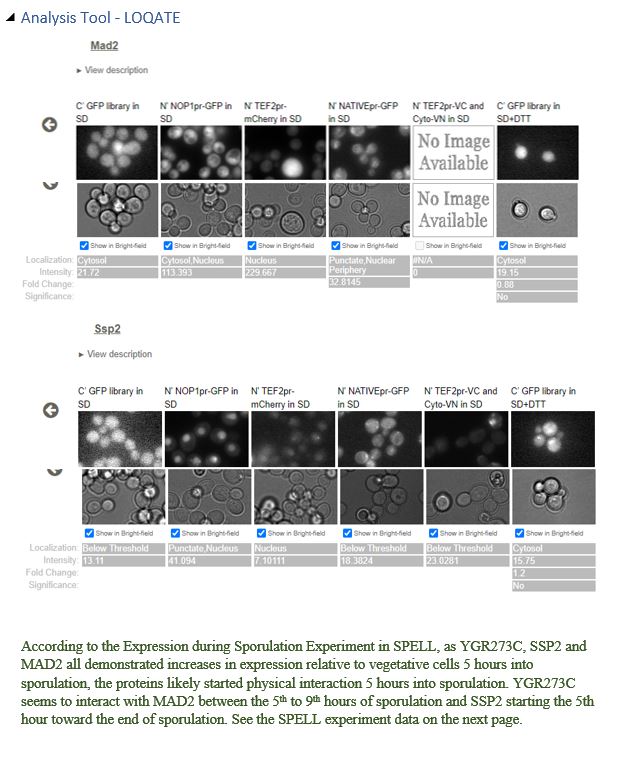
YGR273C is likely localized to the cytosol. YGR273C physically interacts with SSP2 and MAD2. For SSP2, LoQate shows that it punctuates through the nuclear membrane and exists in the nucleus and cytosol with GFP stain. However, the SGD database describes SSP2 as localized to the spore wall. It’s possible that SSP2 eventually settles to the spore wall at a later stage of sporulation. For MAD2, LoQate shows that it exists mainly in the nucleus but is also visible with GFP stain in the cytosol. Since YGR273C is not in the nucleus by NucPred prediction and is likely cytoplasmic by Port II prediction, cytosol becomes a logical place for YGR273C to contact SSP2 and MAD2 physically. Click here to see localization results from various tools.
From SPELL data, YGR273C shares a similar expression pattern over time with proteins involved in sporulation and spore wall formation, such as SPS22, LDS1, SPO19, SPR3, and CDA1. Exploring multiple sporulation expression experiments reveal that these proteins generally start to show elevated expression compared to vegetative growth expression 5-to-8 hours into sporulation, suggesting that YGR273C is more actively involved during the later stage of sporulation, which corresponds to the onset of spore wall formation. Click here to see SPELL search results. Click here to see results from various tools used to predict molecular functions.
Even though the exact molecular functions of YGR273C cannot be established with existing data and tools, it likely participates in sporulation, particularly during the spore wall formation stage. Experiments around determining the relationship between spore wall formation and YGR273C are worth exploring to provide more insights into YGR273C’s functionalities.
Localization
YGR273C is likely localized to the cytosol, the portion of the cytoplasm that is not contained in the organelles. See Figure 1 for the different loci in a Saccharomyces Cerevisiae cell as adapted by the Loqate gene localization prediction tool. See Table 1 for a summary of localization predictions tools used.
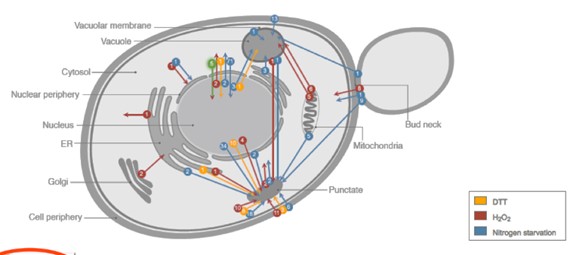
Figure 1. Loci of Saccharomyces Cerevisiae. Source: http://www.weizmann.ac.il/molgen/loqate/help
| Tool Name | Localization Prediction for YGR273C |
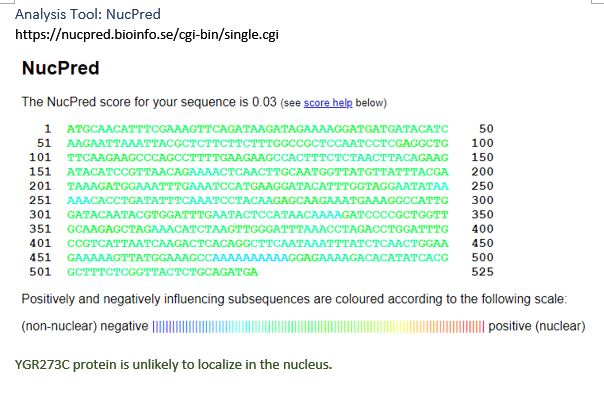
| NucPred | not nucleus |
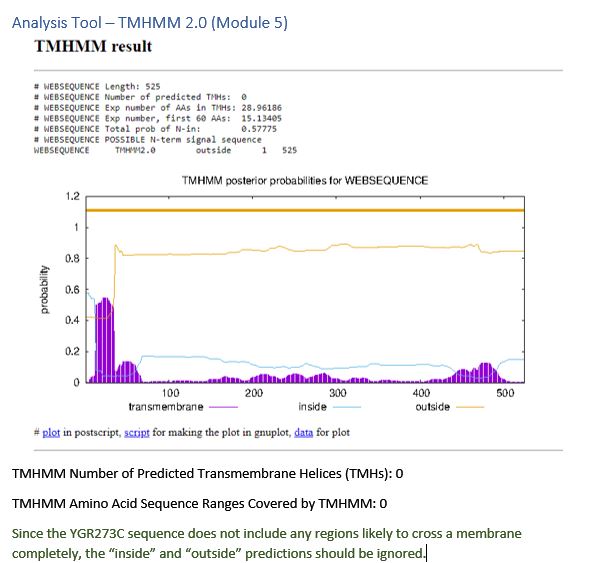
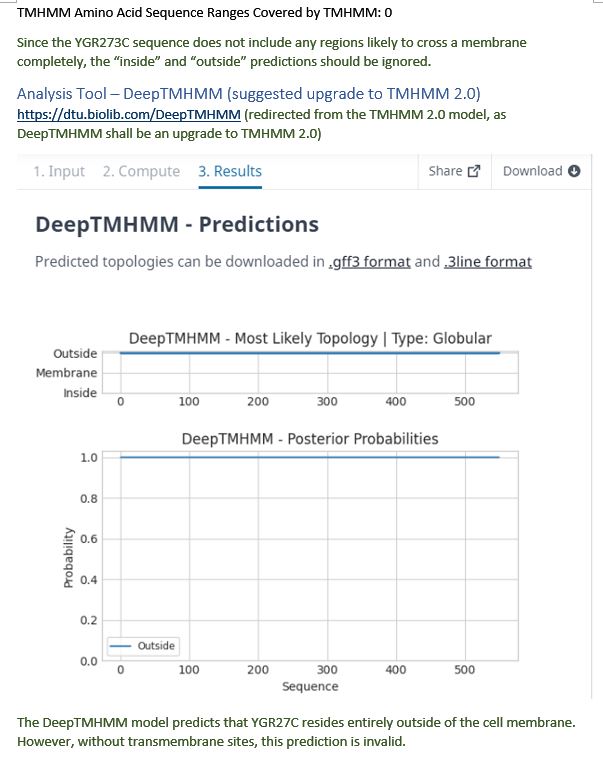
| TMHMM 2.0 | Invalid prediction |
| DeepTMHMM | Invalid prediction |
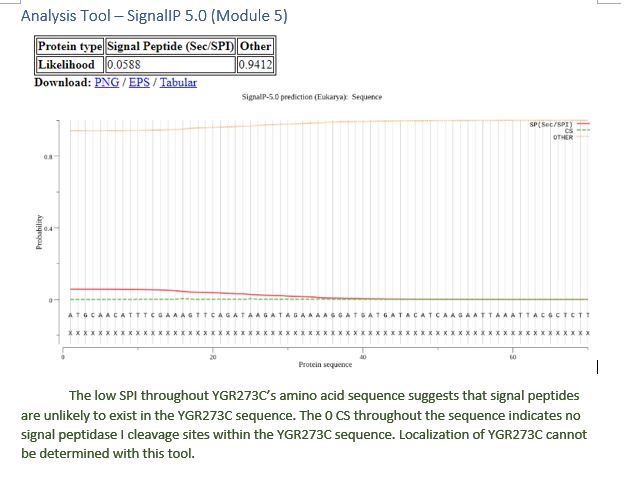
| SignalIP 5.0 | Cannot be determined |
| PortII | 65.2% cytoplasmic |
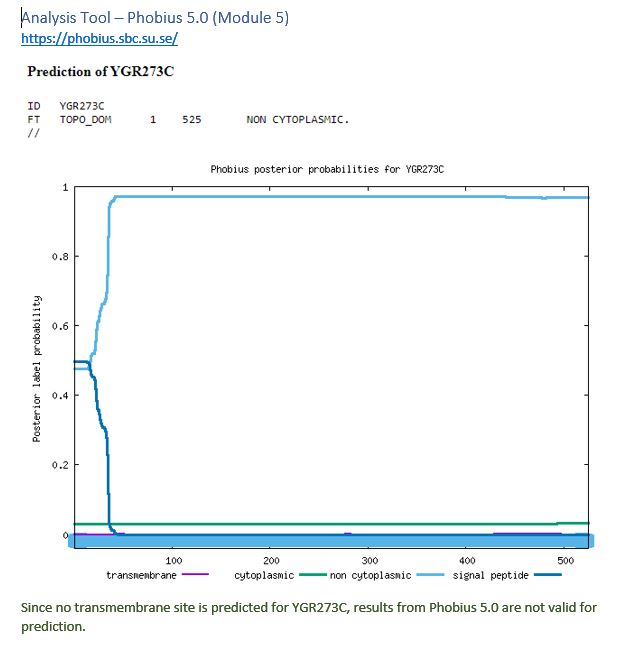
| Phobius 5.0 | Invalid prediction |
| LoQate | no result |
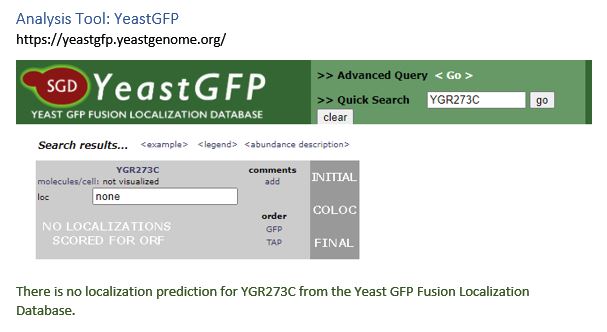
| YeastGFP | no result |
Most of the tools used to predict YGR273C yield inclusive results. NucPred concludes that YGR273C locates outside the nucleus, and Port II weakly predicts that YGR273C is cytoplasmic. To gain better insights into YGR273C’s localization, SSP2 and MAD2, two genes that physically interact with YGR273C, are examined. For SSP2, LoQate shows that it punctuates through the nuclear membrane and exists in the nucleus and cytosol. SGD database describes SSP2 as localized to the spore wall. For MAD2, LoQate shows that it exists mainly in the nucleus but is also visible with GFP stain in the cytosol. Since we learn that YGR273C is not in the nucleus by NucPred and is likely cytoplasmic, according to Port II, cytosol becomes a logical place for YGR273C to contact SSP2 and MAD2 physically. Click on the images below to see interaction results and Loqate results for SSP2 and MAD2.
Molecular Function
The exact molecular functions of YGR273C cannot be determined with current data and tools. However, results from various tools suggest that YGR273C is likely involved in the spore formation process, providing non-essential add-on value to spore wall formation. See Table 2 for a summary of the prediction tools used and their corresponding result.
| Tool Name | Result |
| NCBI CDD Search | N/A |
| PFAM | PFAM DUF2406 also uncharacterized |
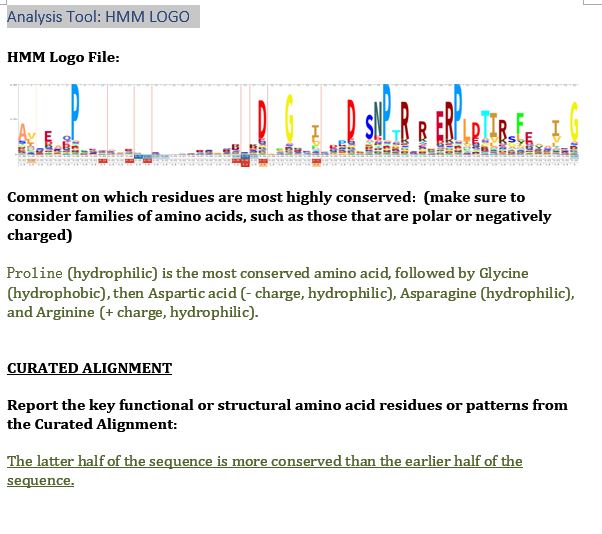
| HMM Logo | Proline (hydrophilic) is the most conserved amino acid, followed by Glycine (hydrophobic), then Aspartic acid (- charge, hydrophilic), Asparagine (hydrophilic), and Arginine (+ charge, hydrophilic). The latter half of the sequence is more conserved than the earlier half of the sequence. |
| Protein Data Bank | N/A |
| SUPERFAMILY | N/A |
| SMART | N/A |
| GENE3D/CATH | N/A |
| Panther | Matches MEIOTICALLY UP-REGULATED GENE 9 PROTEIN (PTHR28186), with no data on protein class and notes on gene ontologies |
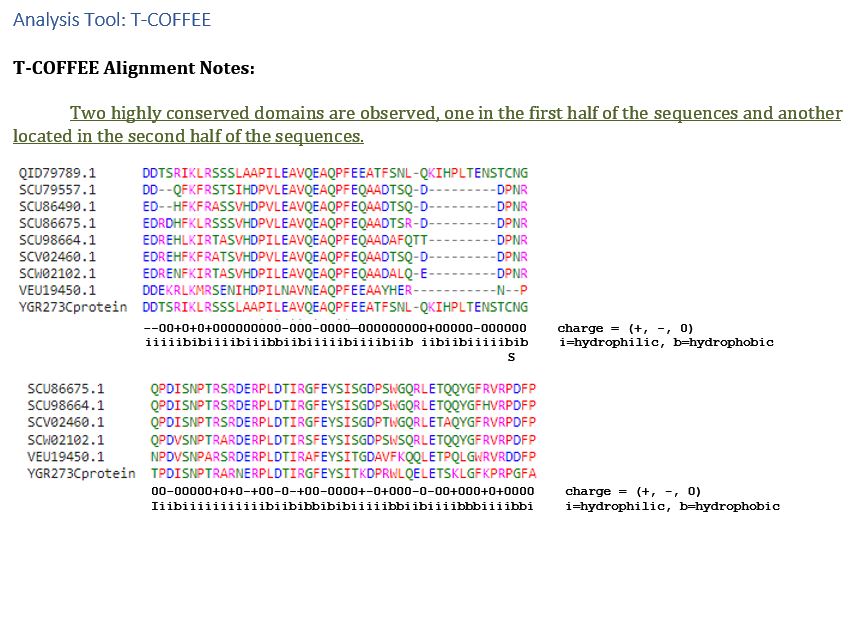
| T-COFFEE | Two highly conserved domains are observed, one in the first half of the sequences and another located in the second half of the sequences. However, none of the proteins compared have characterized functionalities. |
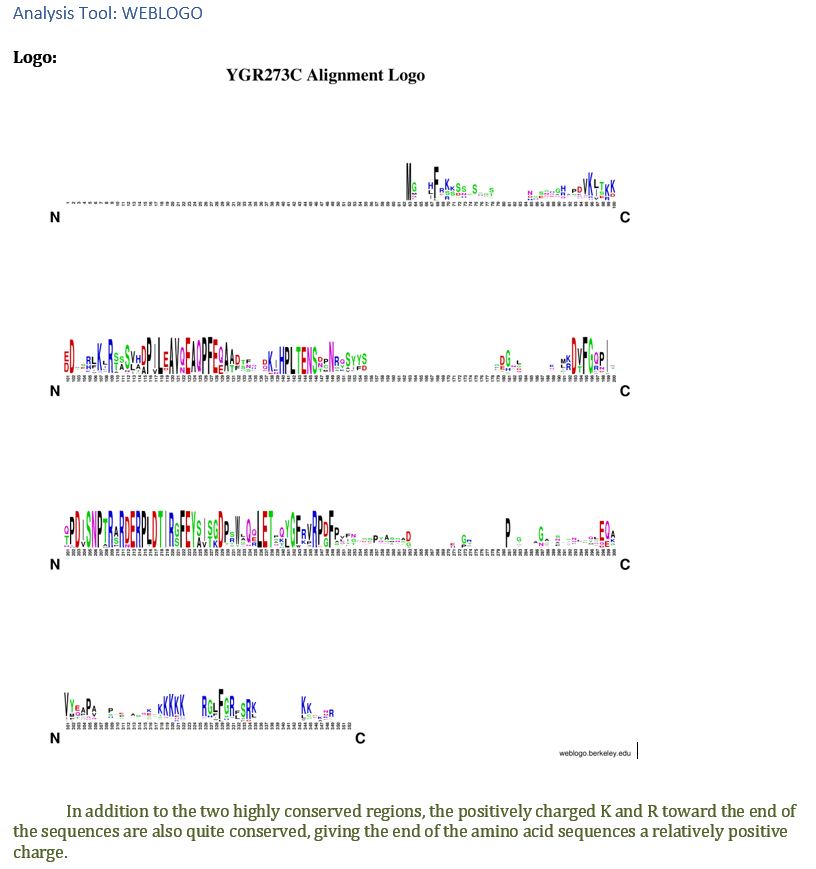
| WEBLOGO | the positively charged Lysine(K) and Arginine(R) amino acids toward the end of the sequences are also quite conserved, giving the end of the amino acid sequences a relatively positive charge. |